Connecting biophysics trainees across Canada!












Université de Montréal, Montreal, QC

Career Speaker Presentations

Career Speaker Panel Discussion
Networking Session
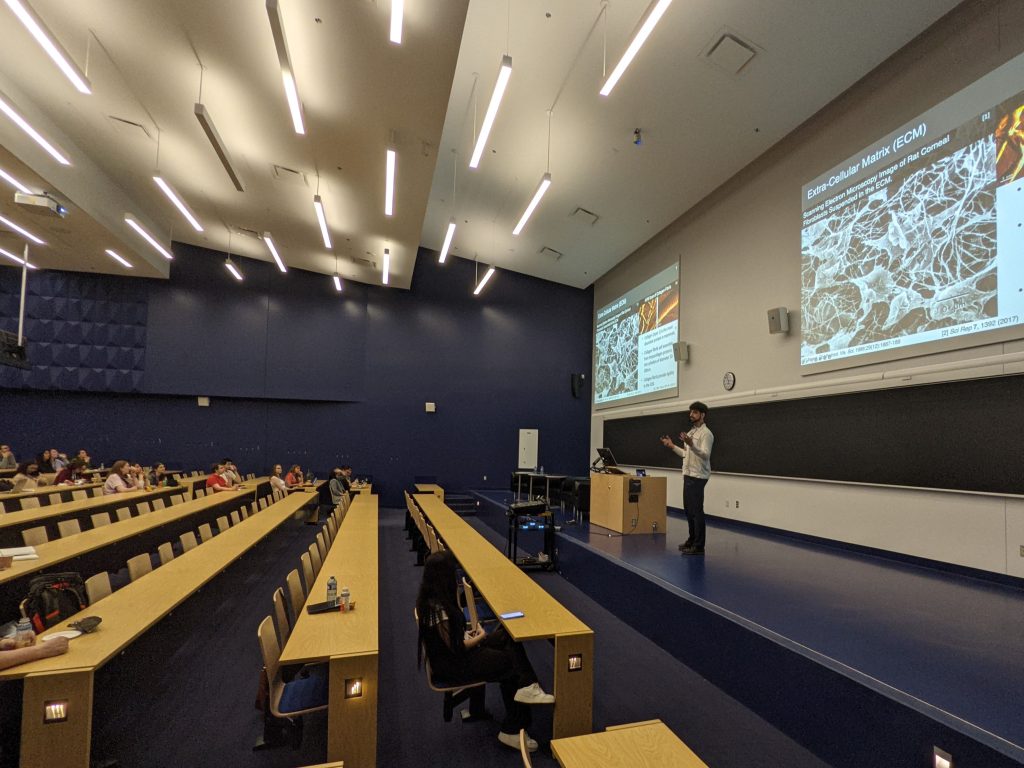

Trainee Presentations
The Trainee Symposium marked the commencement of the 9th annual Biophysical Society of Canada Meeting bringing together trainees from coast to coast. It provided attendees with valuable opportunities to present their research, engage with leaders in various scientific fields, and participate in enriching networking opportunities. Thank you to all our invited and contributed speakers for making this event a great success.
Virtual


During Biophysics week, we heard from Dr. John Dutcher about his experience in transforming a serendipitous discovery of a unique nanoparticle in his lab into a biotechnology company. We also followed along with Sara Evans as she led a gastronomy cooking class on jellies while teaching us the underlying science and showing us neat experiments we can do in our own home.
See Sara’s gastronomy cooking class event highlighted in the Biophysical Society Newsletter here.
must be signed in to X to view

The Biophysical Society of Canada has been connecting biophysicists across Canada since 1985 through annual meetings, events, awards and programs to develop, grow and enrich Canadian biophysics research.
For more information or questions about the BSC, please contact Claudiu Gradinaru.
For website related questions, please contact Isaac Li.